|
Cold Spring Harbor: Cold Spring Harbor Laboratory Press. Removal of 3´ overhangs to form blunt ends Initially, the Klenow fragment of DNA-dependent DNA polymerase I (involved in replication and repair in the bacterium Escherichia coli ), was employed in PCR amplification Saiki et al., 1985 Mullis et al., 1992 Saiki et al., 1986. 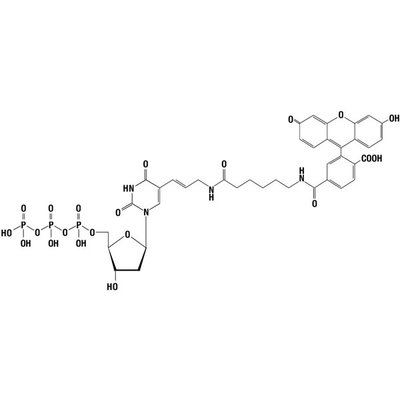
Fill-in of 5´ overhangs to form blunt ends.DNA sequencing by the Sanger dideoxy method.The other small fragment has 5'→ 3' exonuclease activity, but lacks 5'→ 3' polymerase activity and 3'→ 5' exonuclease activity of klenow fragment. It has 3'→ 5' exonuclease activity for proof reading activity Pol I can be cleaved by mild treatment with subtilisin into two fragments the larger fragment is known as the Klenow fragment. This klenow fragment lacks 5'→ 3' exonuclease activity. 10.1101/pdb.top100743 Abstract Escherichia coliDNA Pol I can carry out three enzymatic reactions: It possesses 5' 3' DNA polymerase activity and 3' 5' and 5' 3' exonuclease activity. 1 kb DNA Ladder is stable for at least 3 months at 4C. 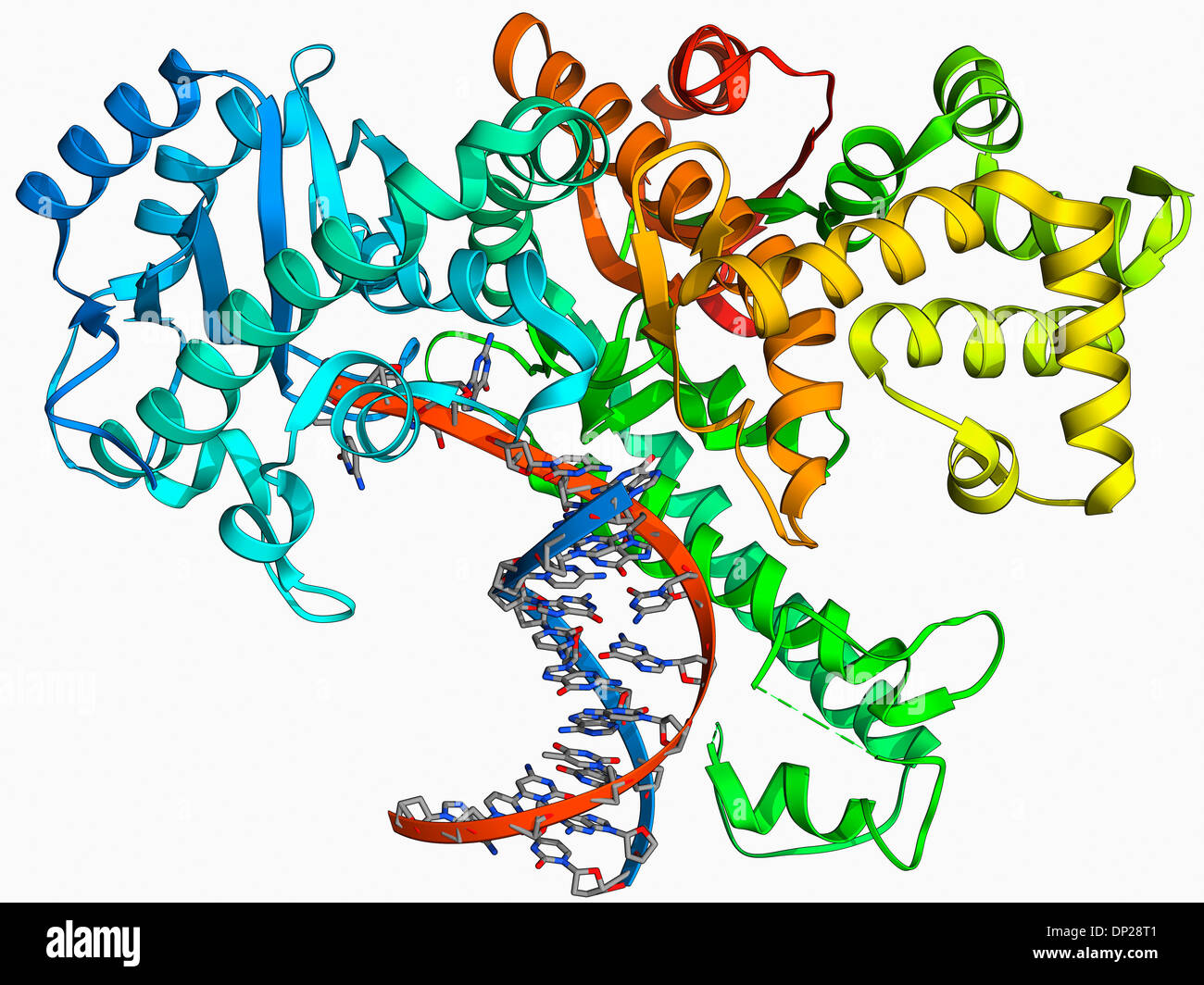
Use -32 P dATP or -32 P dTTP for the fill-in reaction. On the left side, upon treatment with protease, two fragments are formed, a large fragment called the klenow fragments and a small fragment with 5'→ 3' exonuclease activity. coli DNA polymerase I Klenow fragment, Bst polymerase, Phi-29 polymerase, Bacillus subtilis Pol I (Bsu), as well as single-stranded DNA binding proteins from E. All fragments have a 4-base, 5 overhangs that can be end labeled using T4 Polynucleotide Kinase (M0201) or filled-in using DNA Polymerase I, Klenow Fragment (M0210) (1). On the right side is the DNA Polymerase I before treating with protease. Refer this figure for clear understanding.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
